Deep Mining Tools for LncRNA Data: RIP Technology
Long non-coding RNA (lncRNA) refers to RNA molecules longer than 200 nucleotides that typically do not encode proteins. Instead, they function as RNA molecules in various biological activities, such as epigenetic regulation, cell cycle regulation, and cell differentiation regulation, becoming a hot topic in genetics research.

Figure 1 Mechanisms of lncRNA action
Generally, the mainline of lncRNA functional research includes three major steps:
High-throughput screening: Samples are grouped and subjected to high-throughput sequencing techniques (such as RNA-Seq) and specific databases for high-throughput screening. By evaluating the coding potential of transcripts, it can be determined whether they have coding potential, thereby identifying whether a transcript is an lncRNA.
Identification of candidate lncRNAs: Genes or lncRNAs that are differentially expressed between different samples (or sample groups) are identified to obtain candidate RNAs with significant differential expression. Among the non-coding RNAs obtained from sequencing, those with large fold changes, high expression levels, and relevance to the research direction are usually prioritized.
Functional analysis and validation: Based on bioinformatics predictions, experiments are designed to validate the biological functions of non-coding RNAs. For example, real-time quantitative PCR and in situ hybridization techniques can be used to verify the expression of RNA under specific samples or conditions. Additionally, gene knockdown/knockout (using siRNA, shRNA, or CRISPR/Cas9 to reduce lncRNA expression levels) or overexpression experiments (using overexpression vectors to increase lncRNA expression levels) can be conducted to observe phenotypic changes. To delve deeper, techniques such as RNA pull-down, RIP, and dual-luciferase assays can be employed to detect proteins or RNAs interacting with the target RNA, further elucidating the biological functions of RNA. Subsequently, the functions and regulatory mechanisms of RNA can be validated in animal or cell models to confirm their importance under physiological and pathological conditions.
A thorough understanding of lncRNA mechanisms can help identify potential disease mechanisms and aid in the search for new therapeutic targets. In particular, many drugs achieve their therapeutic effects by interfering with specific RNA-protein interactions.
RIP and RNA pull-down are key tools for in-depth study of RNA-protein interactions. RIP uses specific antibodies against target proteins to isolate and purify RNA-protein complexes, while RNA pull-down captures interacting proteins using RNA probes. Today, we will introduce the RIP technique, which is described as follows:
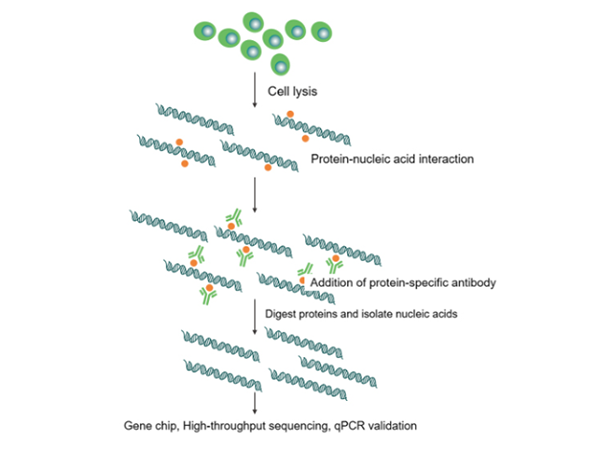
RIP technology (RNA Binding Protein Immunoprecipitation) is a technique for studying RNA-protein interactions within cells and is a powerful tool for understanding the dynamic processes of post-transcriptional regulatory networks. This technique primarily uses specific antibodies against target proteins to precipitate RNA-protein complexes. Subsequently, the RNA bound to the complexes is isolated and purified for validation by q-PCR or high-throughput sequencing analysis, mainly for QPCR validation of interactions between known proteins and known RNAs, and for screening unknown RNAs interacting with known proteins using next-generation sequencing.

Figure 3 Principle of RIP technology
Common Questions:
1. Can RIP be used to separate nuclear and cytoplasmic RNA-protein complexes?
Theoretically, protein extraction kits that can separately extract nuclear and cytoplasmic proteins can be used to separate samples before conducting RIP experiments. However, the reagents in the kit must not contain RNase or DNase, and they must be compatible with downstream antibody-bead incubation.
2. The concentration of RNA pulled down from the sample is too low. How can this be improved?
Possible reasons include insufficient sample volume, consider increasing the initial sample amount, or incomplete lysis, such as insufficient grinding of tissue samples.
3. Abnormal melting curve
Non-specific amplification or primer dimers may occur; redesign primers.
4. There is no difference in Ct values between IP and IgG samples. What could be the cause?
It may be due to the antibody not enriching RNA; try a different antibody. Alternatively, high IgG background may be the issue; increase the number of washes or reduce the amount of RNA input in the immunoprecipitation step.
5. How to determine if the RIP experiment is successful?
Detection of target protein signals in both IP and input groups by WB after RIP indicates a successful experiment.
|
Catalog No. |
Product Name |
Specifications |
|
RNA Pull Down Kit |
6T |
|
|
DNA Pull Down Kit (Plant) |
6T |
|
|
DNA Pull Down Kit (Animal) |
6T |
|
|
RNA Immunoprecipitation (RIP) Kit |
6T | |
|
Chromatin Immunoprecipitation (ChIP) Kit |
22T |
|
|
Immunoprecipitation (IP/CoIP) kit |
50T |
|
|
Protein A/G Magnetic IP/Co-IP Kit |
10T/50T |
AntBio provides antibodies, proteins, ELISA kits, cell culture, detection kits, and other research reagents. If you have any product needs, please contact us.




